What Are the Stages of Medical Software Development?
The development of medical software involves several fundamental stages. According to the IEC 62304 standard, the software life cycle includes planning, requirements analysis, design, implementation, verification, validation, and system maintenance. From a developer’s perspective, the implementation phase can be further divided into preprocessing, neural network implementation, and postprocessing.
In addition, the standard defines processes for risk management, configuration management, and problem resolution. This means that medical software development is not limited to just coding and testing. Every stage must be thoroughly documented and aligned with safety and regulatory requirements.
Requirements Analysis and System Architecture Design
In medical software development, requirements analysis should first clearly define the purpose of the system. In the case of medical image segmentation, this means specifying which diagnostic problem the system is intended to solve. Examples include automatic segmentation of tumors in MRI scans (as shown in Figure 1) or delineation of anatomical structures in CT images.
It is also important to define the expected level of accuracy and acceptable error tolerance. Furthermore, the requirements must consider how the results will be presented to the system’s end users, such as clinicians. This ensures that subsequent design decisions are aligned with the system’s real clinical application.
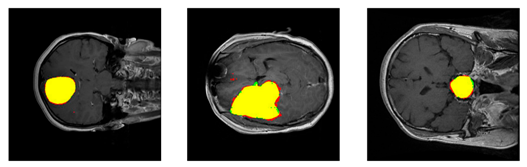
Figure 1. Examples of segmentation results for meningioma, glioma, and pituitary tumors, respectively. Figure sourced from [1].
Preprocessing
Before training a model, medical data must be properly prepared. Preprocessing includes operations that enhance data quality and standardize input, making it suitable for subsequent stages. A common operation in semantic segmentation preprocessing is z-score normalization. This involves subtracting the mean intensity of the image voxels and dividing by the standard deviation, transforming the data to have a mean of 0 and a standard deviation of 1. This approach reduces the impact of differences between training datasets and facilitates model generalization to data from new sources.
Another key component of preprocessing is data augmentation, which increases the diversity of the training set. In medical segmentation, common augmentations include geometric transformations such as rotations, scaling, and flipping. Intensity augmentations are also frequently applied, including contrast adjustment, brightness modification, and noise simulation.
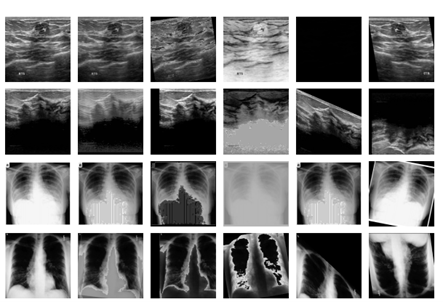
Figure 2. Examples of augmented medical images. Figure sourced from [2].
Neural Network Implementation and Training
During the implementation phase, the architecture of the neural network is coded and the model training procedure is defined. In practice, libraries such as PyTorch or TensorFlow are used, providing flexible tools to build deep learning models. Increasingly, architectures are not designed from scratch; instead, predefined and validated frameworks are applied. A good example is nnU-Net [3], a framework specifically developed for medical image segmentation. nnU-Net automatically adapts the network architecture, training parameters, and preprocessing steps to a given dataset. This reduces the need for manual hyperparameter tuning and lowers the risk of implementation errors. A competitive alternative to nnU-Net is U-Mamba [4], which has been described in one of our previous posts: Mamba Rising: Are State Space Models like U-Mamba Going to Replace Ordinary U-Net?
In medical software development, neural network training often relies on cloud infrastructure, such as AWS, to meet high computational and memory demands. The cloud allows flexible allocation of GPU resources on demand, scaling resources according to dataset size, and parallel training of multiple models. This makes the training process faster and more reproducible.
Postprocessing
After obtaining the raw outputs from a segmentation model, postprocessing is performed to improve the quality and clinical reliability of the masks. Models such as nnU-Net typically return class probability maps for each voxel in float format (e.g., 0–1), where the value indicates the likelihood of belonging to the target class. To obtain the final binary mask, these maps are subjected to thresholding, which converts the probability map into a discrete mask. A common threshold is 0.5, meaning a voxel is assigned to the positive class if its probability exceeds this value. In practice, the threshold can be optimized for a specific task.
After thresholding, morphological spatial operations are often applied to improve the continuity of anatomical structures and remove artifacts. Examples include removing isolated small regions and filling holes within larger segments. Operations such as opening and closing help smooth edges and stabilize the masks.
More advanced approaches use unsupervised postprocessing networks, such as autoencoder models trained to reconstruct anatomically plausible masks from raw predictions. These methods learn the space of valid masks and project the segmentations back into that space, improving both visual quality and anatomical consistency. An example of such a method is described in [5].
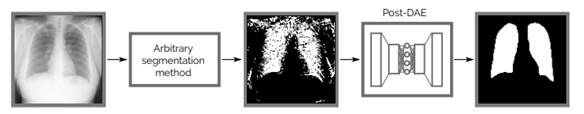
Figure 3. Example of using an autoencoder network in postprocessing. Figure sourced from [5].
Deployment and System Maintenance
The final deployment of medical software involves installing the application for end users, integrating it with existing systems, and monitoring its performance in the production environment. It is essential to provide a scalable and secure environment and implement mechanisms for automated updates. After deployment, the system requires continuous maintenance – technical support includes tracking bugs, responding to failures, and collecting user feedback. In summary, deployment is not the end of the process. The system must be continuously maintained and monitored for both effectiveness and regulatory compliance. In practice, this means constantly adapting and improving the software to meet clinical and technical requirements over the long term.
Resources
[1] Francisco Javier Díaz-Pernas, Mario Martínez-Zarzuela , Míriam Antón-Rodríguez, David González-Ortega. (2021). A Deep Learning Approach for Brain Tumor Classification and Segmentation Using a Multiscale Convolutional Neural Network.
[2] Zhaoshan Liua, Qiujie Lvb, Yifan Lia, Ziduo Yanga,c, Lei Shena. (2024). MedAugment: Universal Automatic Data Augmentation Plug-in for Medical Image Analysis.
[3] Fabian Isensee, Jens Petersen, Andre Klein, David Zimmerer, Paul F. Jaeger, Simon Kohl, Jakob Wasserthal, Gregor Koehler, Tobias Norajitra, Sebastian Wirkert, Klaus H. Maier-Hein. (2018). nnU-Net: Self-adapting Framework for U-Net-Based Medical Image Segmentation.
[4] Jun Ma, Feifei Li, Bo Wang. (2024). U-Mamba: Enhancing Long-range Dependency for Biomedical Image Segmentation.
[5] Agostina J. Larrazabal, Cesar Martinez, Enzo Ferrante. (2019). Anatomical Priors for Image Segmentation via Post-Processing with Denoising Autoencoders.